Enhanced Compound Identification for Quad Based GC-MS with the TAMI Software
Pushing sample identification by GC-MS to its limits
Aviv Amirav and Tal Alon (June 2022)
In Short
Electron Ionization (EI) mass spectra contain a vast amount of information that can be considered during an identification procedure. It might be surprising to know that most of us examine and utilize only a part of that information and as a result decrease the probability of a correct identification and in many cases cannot identify the analytes at all. It might also be surprising to know that it is rather easy to consider more of the data and improve identification results.
When we analyze an EI mass spectrum we usually relay on library identification, which provides us the most resembling spectra from the library (with almost no effort on our part). Library identification is an important tool but it has its shortcomings, and it often fails without us knowing that it did.
We have developed new and improved compound identification methods and software which utilize the extra informative data found in EI spectra but ignored by library identification algorithms. These methods which rely on a combination of Isotope Abundance Analysis (IAA), unit resolution mass accuracy and library search were incorporated within a software we call TAMI (Tal Aviv Molecule Identifier).
The TAMI software provides the following benefits for "pushing sample identification by GC-MS to its limits":
A) Automatically confirms or rejects NIST library identification for improved confidence level in the library identification.
B) Provides accurate elemental formula from low cost unit resolution quadrupole GC-MS data.
C) Provides the above with standard centroid data files while using standard data analysis software such as Chemstation or MassHunter or Chromeleon.
D) Very easy to use with one click operation.
A Brief Explanation on IAA
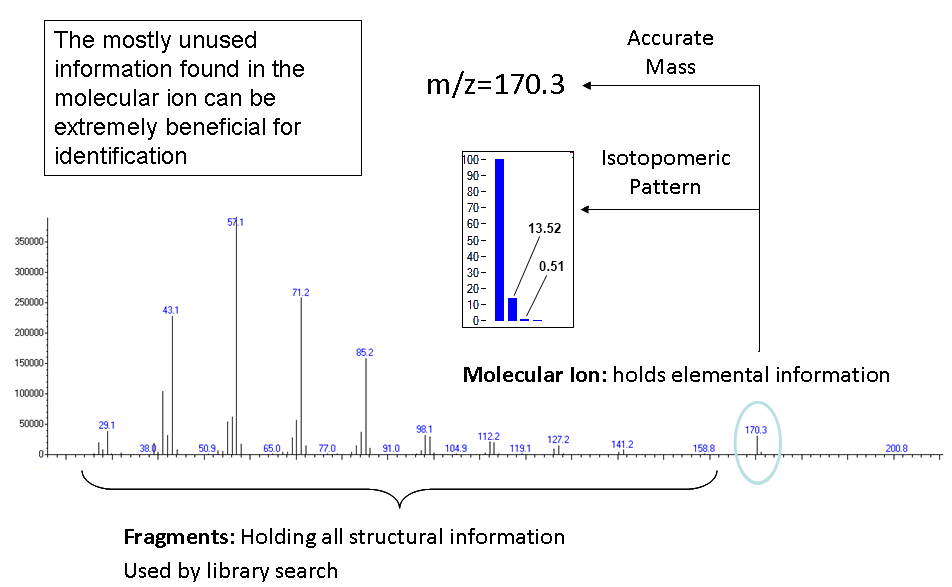
In most cases the molecular ion and each fragment are represented as a
collection of masses and not as a single mass. The reason for this is the
existence of the isotopes as can be seen in the figure below.
These ions which belong to the same elemental composition are called
“isotopomers” and their relative abundances are attributed to the natural
abundances of the isotopes in nature. The abundances of these isotopomers can be
used to generate the elemental formula, providing results which are similar to
those provided by costly high resolution instruments.
A Brief Explanation on Mass Accuracy Filtering
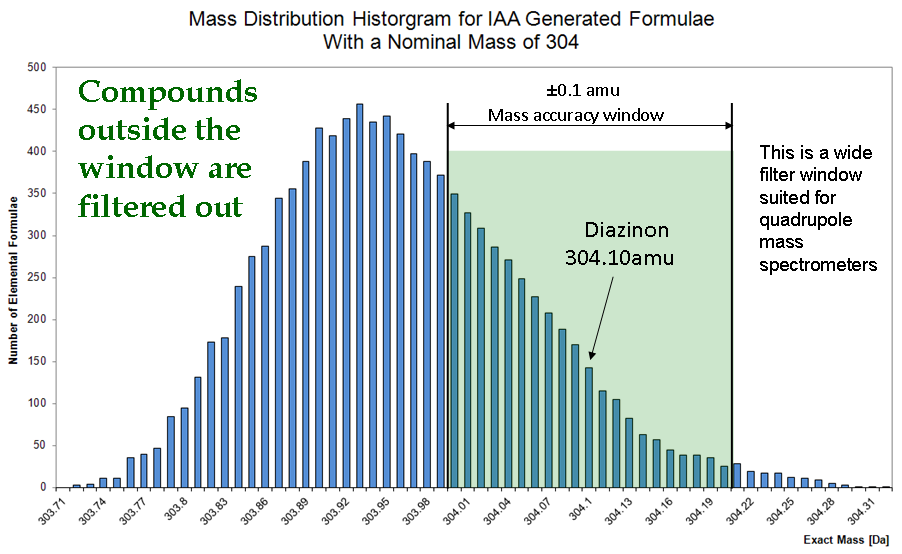
Standard Quadrupole Centroid files usually provide masses with errors under 0.14amu, and this rough measurement is still very beneficial as it helps TAMI’s identification procedure by filtering out impossible elemental compositions with masses outside this ±0.15amu error as can be seen in the following figure.
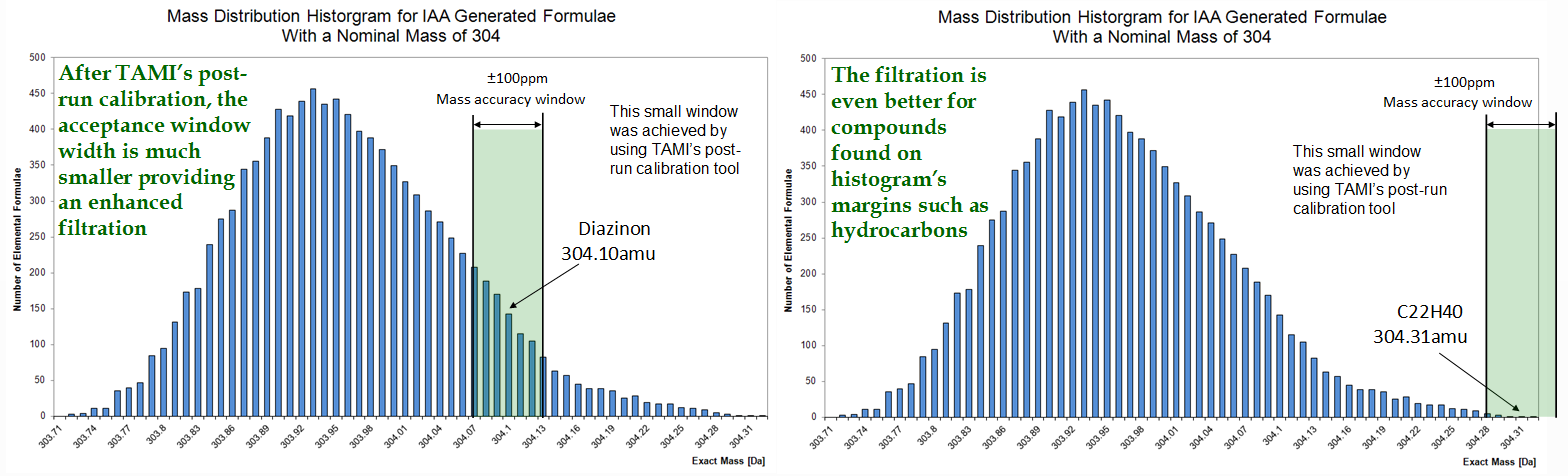
It is however possible, with a properly tune MS and a good centroiding algorithm to bring quadrupole mass measurements under ±100ppm error. This reduced window size dramatically improves identification results as can be seen in the following figure.
Library - IAA Algorithm
TAMI automatically links with the MS data analysis software (such as Agilent’s Chemstation or MassHunter) and with the NIST Mass Spectral Library. Whenever a mass spectrum is sent to be analyzed by the NIST library TAMI grabs a copy of the spectrum and can then apply the new analytical methods:
TAMI analyzes both the spectrum’s isotopomeric patterns and NIST’s results, two physically orthogonal non-related sets of data, and produces a report which clearly supports the library results or rejects them. The identification probability is recalculated and becomes more accurate. Here are two general examples showing how TAMI’s involvement helps with library identification:
Supporting Correct Library Identification : The library can correctly identify a compound but give it a low identification probability due to the similarity of the measured spectrum to other compounds [1]. TAMI’s algorithm will automatically detect that the identification is correct (using IAA on the molecular ion group) and can improve its identification probability.
Rejecting False Library Identification : The library can falsely identify a compound and even give it a decent identification probability, TAMI’s algorithm will automatically detect that the identification is wrong (using IAA) and drastically reduce its identification probability. TAMI might also indicate that the compound which is number 2 on the library hit-list is in fact more likely to be the measured compound. in such a case TAMI can apply another powerful independent algorithm to identify the compound as described below.
Independent Algorithm : IAA + Mass Accuracy Filter
In cases of library failure, and there can be several reasons for such a failure which is more common than most people realize, TAMI can apply an independent analysis method which starts with the automatic identification of the molecular ion in the measured spectrum, and continues with full isotope abundance analysis of the molecular pattern and mass accuracy analysis (filter). The result of such analysis is a “hit-list” of elemental compositions sorted according to their probability.
The previous example showed how TAMI identifies a false library identification, and as can be seen in the next figure TAMI provide that extra step and identifies the compound independently.
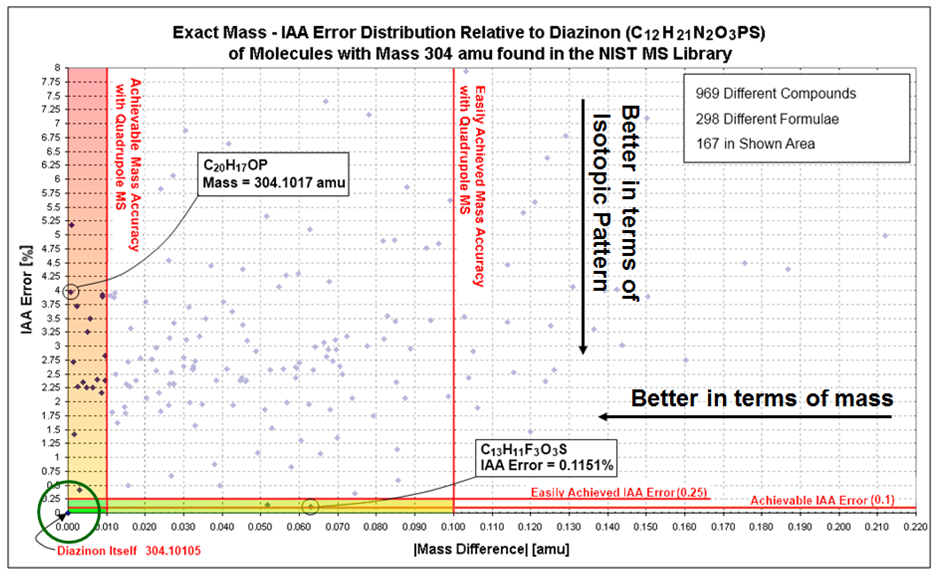
This algorithm combines two distinct physical characteristics which are linked with the elemental composition of the compound - isotope abundance analysis and measured mass. The figure below show how this combination works, where each axis is completely independent of the other - leaving only a few possible elemental compositions and one which is most probable.
For the best results the mass spectrometer used should be calibrated, or else TAMI can calibrate it using any raw (profile) data file which includes a known compound. This calibration reduces the mass error to under ±100 ppm which leads to better identification.
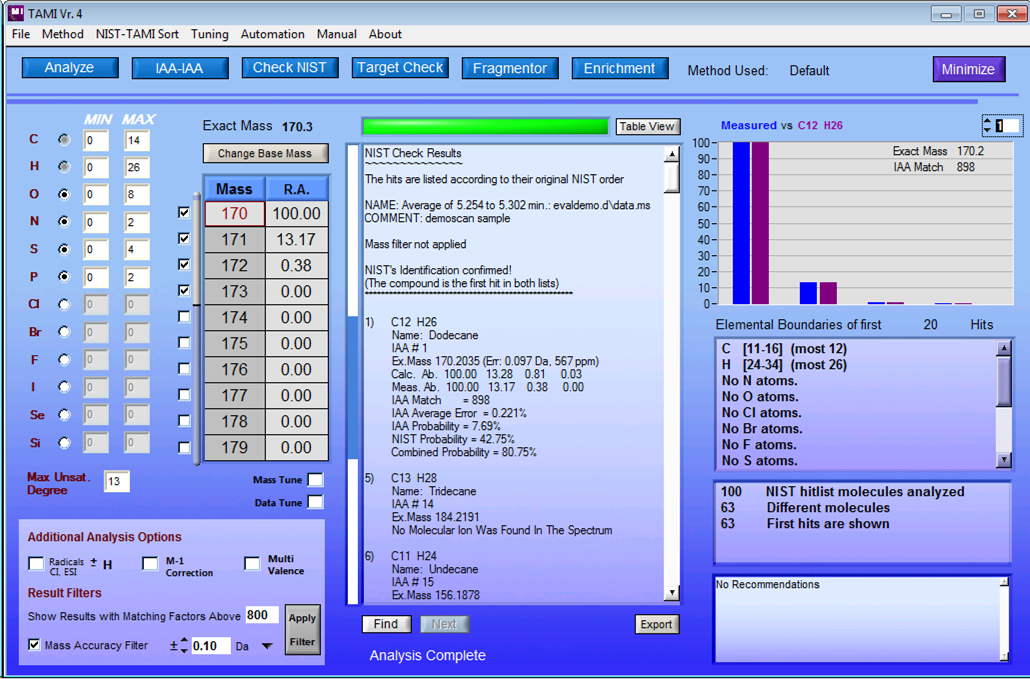
TAMI Main Software Window and Demonstration Video
Further Information
Tal Alon and Aviv Amirav "Isotope Abundance Analysis Method and Software for Improved Sample Identification with the Supersonic GC-MS” , Rapid Commun. Mass Spectrom. 20, 2579-2588 (2006).
Tal Alon and Aviv Amirav "A Comparison of Isotope Abundance Analysis and Accurate Mass Analysis in their Ability to Provide Elemental Formula Information" J. Am. Soc. Mass Spectrom. 32, 929-935 (2021).
Tal Alon and Aviv Amirav "Enhancing the Identification Capabilities of EI GC-MS - How Quadrupole GC-MS can compete with High Resolution TOF” , Advanced GC-MS Blog Journal (2013).
Further information on the TAMI software can be found in its Aviv Analytical TAMI website